Improving Breast Cancer Detection With Upsampling And Resizing Algorithms In Image Processing
Abstract—Our study focuses on the detection of breast cancer using medical image analysis. The researchers explore the effectiveness of various oversampling methods in improving the performance of deep learning models for breast cancer detection. The dataset used in the study has a severe class imbalance with a disproportionate number of cancerous and non-cancerous examples. Six oversampling methods are evaluated in this study. Each oversampling method is applied to the dataset, and the augmented data is used to train deep learning models. The performance of each oversampling method is evaluated using metrics such as accuracy, precision, recall, and F1-score. The results demonstrate that oversampling methods significantly enhance the performance of deep learning models for breast cancer detection. SVM-SMOTE and ADASYN consistently outperform other methods, achieving the highest F1 scores on both ResNet-50 and AlexNet architectures. The findings also suggest that the choice of oversampling method has a substantial impact on model performance, emphasizing the importance of selecting an appropriate oversampling technique for imbalanced data. Overall, this study highlights the significance of addressing class imbalance in medical image analysis and provides valuable insights into the effectiveness of different oversampling methods in improving the performance of deep learning models for breast cancer detection.
Keywords—deep learning, breast cancer, image preprocessing, resizing, imbalanced dataset, convolutional neural networks.
I. INTRODUCTION
Breast cancer is a type of cancer that starts in breast cells and has the second-highest incidence worldwide after lung cancer [1]. In 2015, there were an estimated 17.5 million cancer cases and 8.7 million deaths. The risk of breast cancer increases with age and can occur in women of any age post-puberty . Early-stage diagnosis significantly improves cure and survival rates, emphasizing the importance of screening and early detection methods. Recently, the use of advanced technologies such as deep learning and artificial intelligence in the medical field has increased rapidly. There has been a rise in studies on cancer detection using deep learning [2]. These techniques assist experts by identifying patterns and complex relationships in large datasets, improving early detection and treatment processes.
In addition to these methods, sampling methods, i.e. techniques for selecting subsets from a given data set to provide a better understanding and analysis of the data, are used to eliminate the problem of unbalanced data sets, which is frequently encountered in machine learning and data analysis projects.
Sampling is used to streamline processing and save time, especially with large data sets. Interpolation techniques can also be used to estimate missing data or to make the data set smoother.
A. Related Studies
The study by Janowczyk and Madabhushi explores the application of deep learning (DL) to image analysis challenges in digital pathology [3]. They highlight the difficulties in analyzing digital pathology images due to slide preparation problems, variations in staining and scanning, and biological differences. However, they show that DL can be successfully employed in this field.
They conducted their experiments using more than 1200 digital pathology images. This comprehensive tutorial discusses the potential benefits of DL in digital pathology image analysis. By leveraging DL approaches and tools like Caffe, researchers and practitioners can improve the accuracy and efficiency of tasks such as diagnosis, counting, segmentation, and classification in the digital pathology context.
In their discussion, Reza, Md Shamim, and Jinwen Ma emphasize the need for early detection of breast cancer, which is a leading cause of cancer-related deaths globally [4]. They highlight the significance of pathologists' expertise in diagnosing breast cancer. The authors suggest that computer-aided diagnostic tools that employ automatic classification of microscopic images can greatly aid clinical experts. Two datasets, one from http://andrewjanowczyk.com/wp-static/IDC_regular_ps50idx5.zip/ and the other from https://web.inf.ufpr.br/vri/databases/, were used for the study. Both datasets suffer from class imbalance, which can impede the performance of convolutional neural networks (CNN) during parameter learning. To address this imbalance issue, various oversampling and undersampling techniques were employed to equalize the classes in the training dataset. This approach aimed to match the performance of CNN-based classifiers with that of a balanced dataset. The study evaluated the results by considering the accuracy of the imbalanced dataset and the integrated zoom-level and image-level accuracy measure. The oversampling technique with replacement achieved the highest recognition rate at the image level (86.13%), while CNN performance on imbalanced data was 85.01%.
CHEN, Yuanqin, et al present an automatic classification method for breast cancer based on a convolutional neural network (CNN) [5]. They introduce a fine-tuned residual network (ResNet) that performs well and reduces training time while automatically extracting features. To address the issue of a small training set, the authors adopt a data augmentation policy to expand the training data, reducing the risk of overfitting. The main contribution of this study is the development of an automatic mammography classification system with high prediction performance, achieved through transfer learning and data augmentation. The experiments are conducted on a public dataset called CBIS-DDSM, which includes 2620 screening film mammography studies. The proposed method exhibits desirable performance in terms of accuracy, specificity, sensitivity, AUC, and loss, achieving 93.15%, 92.17%, 93.83%, 0.95%, and 0.15% respectively. The proposed method demonstrates robustness and generalizability.
This study focuses on improving the accuracy of analyzing medical images for the detection of cancer cells. The researchers plan to implement various preprocessing steps to ensure that important information is not lost during image resizing. The study highlights the importance of preprocessing and addressing data imbalance in order to enhance the performance of deep learning models for cancer detection in real-world applications. A key aspect of the study is the creation of a balanced dataset using a combination of image resizing algorithms and oversampling techniques. The researchers will then compare different deep learning models, including both older and newer approaches, to observe their impact on the accuracy of breast cancer detection using Residual Neural Networks (ResNet-50) and AlexNet models.
II. MATERIAL AND METHODS
A. Dataset
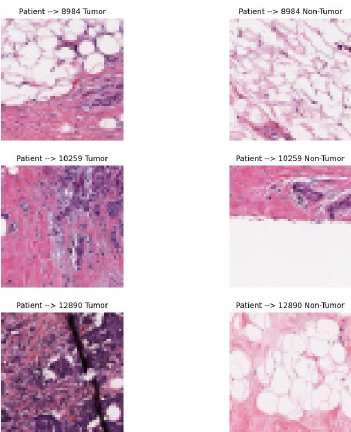
The breast cancer dataset will be used in our study. The dataset was selected as a public dataset on Kaggle. The dataset IDC (Invasive Ductal Carcinoma) consists of 162 full-mount slide images of data samples scanned at 40x. From this, 277,524 50 x 50 patches were extracted (198,738 IDC negative and 78,786 IDC positive). The images are in png format. The file name of each patch is in the following format: u_xX_yY_classC.png, where u is the patient ID (10253_idx5), X is the x-coordinate of where this patch was cropped, Y is the y-coordinate of where this patch was cropped, and C is where 0 is not an IDC and 1 is an IDC. Some of the patches are not 50 x 50. Therefore they will be taken into account when resizing. Some of the extracted 50x50 pixel images can be seen in Fig. 1.
B. Oversampling
Data imbalance is a common problem in machine learning and data analysis projects.It occurs in data sets where the representation of classes or categories is unbalanced. For example, a dataset may have a large difference between the number of cancerous and non-cancerous cells. This imbalance can negatively affect the performance of the model and make it difficult to accurately predict rare classes. Oversampling and undersampling methods can be used to deal with data imbalance. In our study, due to the small amount of data available, we chose oversampling, which is a method of increasing the amount of data.
To briefly talk about the oversampling methods, we use:
•
SMOTE: Addresses data imbalance by creating new synthetic samples among minority class data. It creates new instances by linear interpolations between existing minority class instances.
𝑋𝑛𝑒𝑤=𝑋𝑖+𝛽(𝑋𝑗−𝑋𝑖) (1)
β is the number between 0 and 1. 𝑋𝑗and 𝑋𝑖 are calculated according to the formula at (1).
•
Borderline-SMOTE: It is a variation of SMOTE and focuses on minority class samples near class boundaries.
•
Kmeans-SMOTE: It divides the minority class samples into clusters using the KMeans algorithm and applies SMOTE within each cluster.
•
SVM-SMOTE: SVM (Support Vector Machines) is used to identify class boundaries and synthetic data is generated only among the minority class instances at the boundary.
•
Random OverSampling: A method that randomly selects and replicates instances of a minority class to address data imbalance. Minority class When there are N instances and the majority class When there are M instances( M>N ). To equalize the number of instances of the minority class to M, it is necessary to create =-K=M-N new instances.
•
ADASYN: It generates synthetic examples like SMOTE, but focuses on the more difficult to learn regions of the minority class.
Degree of Difficulty Calculation:
𝑟𝑖= 𝑀𝑎𝑗𝑜𝑟𝑖𝑡𝑦 𝑐𝑙𝑎𝑠𝑠 𝑠𝑎𝑚𝑝𝑙𝑒𝑠 𝑐𝑜𝑢𝑛𝑡 𝐾𝑁𝑁 𝑜𝑓 𝑋𝑖𝐾 (2)
Here 𝑋𝑖 and 𝑟𝑖 is the degree of difficulty of the minority class example. K is value of number of cluster.
As (2) shows, the degree of difficulty is calculated for each minority class sample. This is based on the proportion of majority class instances among the k neighbors of each minority class instance. According to the degree of difficulty, the number of synthetic samples to be generated for each minority class instance is determined. Synthetic instances are generated similar to SMOTE, but with a different number of instances depending on the degree of difficulty.
C. Interpolation
Interpolation is the process of estimating unknown data points between known data points. This process plays an important role in data analysis, especially in filling in missing data and more precise modeling.
•
Area Interpolation: Is a technique used to estimate the area covered by a set of data points in a given region.
•
Cubic Interpolation: Estimates the curve between two points using a cubic polynomial.
Cubic polynomials are usually expressed as in (3):
𝑆(𝑥)=𝑎+𝑏(𝑥−𝑥𝑖)+𝑐(𝑥−𝑥𝑖)2+𝑑(𝑥−𝑥𝑖)3 (3)
•
Lanczos4: Is a technique for high-quality image resampling that uses a modification of the sinc function. This method considers more data points for more accurate results.
•
Nearest Neighbor Interpolation: Nearest neighbor interpolation estimates an unknown point by the value of the nearest known data point. This method is simple and fast, but can produce truncated and jagged results.
•
Linear Interpolation: Is a method of predicting an unknown point between two known data points using a linear line.
•
Linear Exact Interpolation: Is a special case of linear interpolation. This method assumes that all given points have a strictly linear relationship and estimates the intermediate points according to this linear relationship. Equation can be seen in (4).
𝑦(𝑥)=𝑦0+(𝑦1−𝑦0)∗𝑥−𝑥0𝑥1− 𝑥0 (4)
D. Convolutional Neural Networks
It is one of the deep learning methods and is widely used in image processing and computer vision. CNNs can automatically learn and recognize features in images.
ResNet (Residual Network) is a model that aims to improve performance by increasing the number of layers in deep neural networks. It was developed in 2015 by Kaiming He et al [6]. The main innovation of ResNet is that it makes it possible to train very deep networks using a method called residual learning.
AlexNet is a neural network model that has made great progress in the field of deep learning and computer vision. In 2012, AlexNet won the ILSVRC (ImageNet Large Scale Visual Recognition Challenge) competition, marking a milestone in the field of deep learning [7].
First, we applied different oversampling methods to solve the imbalance problem in our medical image dataset, including SMOTE (Synthetic Minority Over-sampling Technique), ADASYN (Adaptive Synthetic Sampling), and Random Over Sampler. After oversampling, we applied different interpolation methods (e.g., nearest neighbor, bilinear, cubic, lanczos4) on the augmented dataset.
Then, we trained the AlexNet and ResNet-50 models on these augmented datasets and evaluated their performance. Our results allowed us to analyze the impact of different oversampling and interpolation techniques on model performance. These analyses are designed to help both models classify patient and non-patient images more accurately.
III. RESULTS
MRI images of thirty patients were recorded in 50x50 pixel patches in RGB format. However, it was observed that some of the patches had different sizes, so they were resized to 50x50. The images were then resized to be suitable for ResNet-50 and AlexNet. In this context, they were resized to 224x224 pixels for ResNet-50 and 227x227 pixels for AlexNet using the relevant algorithm.
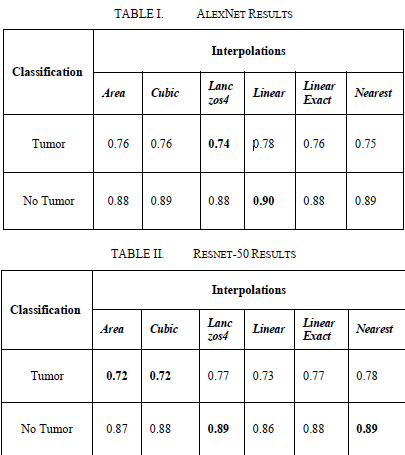
After the above procedures, the results were obtained without using any upsampling method. The results were evaluated separately for AlexNet and ResNet-50. As a result of this evaluation, the classification performance of both models was determined.
Comparisons made according to the F1-Score.
IV. CONCLUSION
According to the results obtained, the classification performance of the AlexNet and ResNet-50 models was analyzed only on interpolated images. In this analysis, particular attention was paid to the cases with a low number of images.
It observed that AlexNet's Lanczos4 interpolation method was more inaccurate in predicting the dwindling number of images. However, in the ResNet-50 model, an F1-Score of 0.72 achieved in the prediction of images with a dwindling number of images using the Cubic interpolation method.
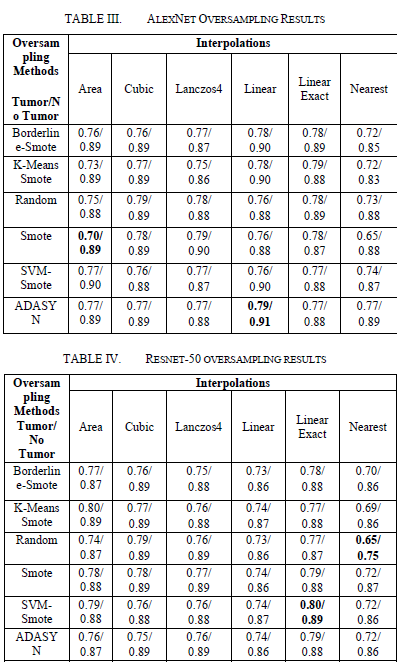
On the AlexNet model, the use of oversampling methods led to an increase in the F1 score of the tumor class. The lowest F1 score observed in the Area/SMOTE combination, while the highest F1 score obtained in the Linear/ADASYN combination. On the ResNet-50 model, the lowest F1-Score recorded with Random Oversampler, while the highest F1 score was obtained with SVM SMOTE.
These results emphasize the effects of oversampling methods on model performance. In the AlexNet model, more advanced oversampling techniques such as Linear/ADASYN observed to provide better performance, while SVM SMOTE achieved the highest performance in the ResNet-50 model.
In future studies, oversampling methods should select, especially if deep learning models are to be used on unstable datasets such as medical imaging datasets. In models such as AlexNet, it is important to choose effective oversampling techniques such as Linear/ADASYN to obtain more balanced and reliable results. Similarly, using more sophisticated oversampling techniques such as SVM SMOTE in models like ResNet-50 can improve model performance and lead to more accurate classification results. These results should be considered in future studies to develop more effective disease detection and classification systems in the field of medical imaging.
REFERENCES
[1]
AZAMJAH, Nasrindokht; SOLTAN-ZADEH, Yasaman; ZAYERI, Farid. Global trend of breast cancer mortality rate: a 25-year study. Asian Pacific journal of cancer prevention: APJCP, 2019, 20.7: 2015.J.
[2]
RAZZAK, Muhammad Imran; NAZ, Saeeda; ZAIB, Ahmad. Deep learning for medical image processing: Overview, challenges and the future. Classification in BioApps: Automation of decision making, 2018, 323-350.
[3]
JANOWCZYK, Andrew; MADABHUSHI, Anant. Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases. Journal of pathology informatics, 2016, 7.1: 29.
[4]
REZA, Md Shamim; MA, Jinwen. Imbalanced histopathological breast cancer image classification with convolutional neural network. In: 2018 14th IEEE International Conference on Signal Processing (ICSP). IEEE, 2018. p. 619-624.
[5]
CHEN, Yuanqin, et al. Fine-tuning ResNet for breast cancer classification from mammography. In: Proceedings of the 2nd International Conference on Healthcare Science and Engineering 2nd. Springer Singapore, 2019. p. 83-96.
[6]
HE, Kaiming, et al. Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition. 2016. p. 770-778.
[7]
KRIZHEVSKY, Alex; SUTSKEVER, Ilya; HINTON, Geoffrey E. Imagenet classification with deep convolutional neural networks. Advances in neural information processing systems, 2012.

